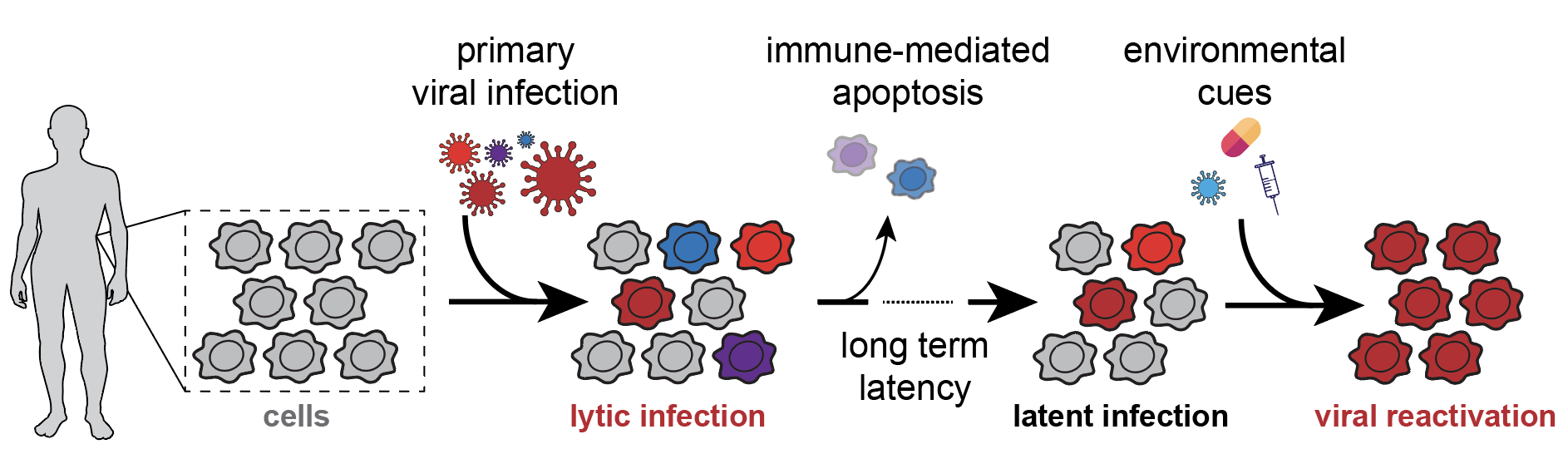
Overview of detection methods
Repo for the main code associated with the quantification of HHV-6 in single-cell data is available here
Repo for the main code associated with the quantification of EBV in whole genomes is available here
Use Serratus to perform a human virus detection screen
For a digest on how to screen all human viruses by all RNA-seq libraries in Serratus, check out the repo here for reproducing the pan-virus screen.
EBV reference genome
The 171,823 nucleotide EBV genome (NC_007605.1) was first
included in December 2013 (hg38 version GCA_000001405.15) as a sink for off-target reads that are often present in
sequencing libraries, to account for pervasive EBV reads present from the immortalization of LCLs (as with the
1000 Genomes Project and related consortia). Importantly, whole genome sequencing (WGS) in the
UK Biobank (UKB) and All of Us (AoU) consortia was performed on whole blood, reflecting that
EBV reads detected would derive from viral DNA from prior infections.
While one can mostly just directly extract rads from the contig, we found highly repetitive regions that were
problematic in finding true viral reads and not background contamination. We recommend removing regions that are
high coverage exceeding an order of magnitude more than the median coverage in the data. For our study, we use these
regions as a quick start for doing this yourself (R code):
wipe_here <- sort(c(36389:36516, 52012:52034, 95997:96037,163596:163617, 36390:36514, 95997:96037))
HHV-6B reference genome
To detect HHV-6 transcripts in existing single-cell data,
we developed a kallisto|bustools workflow to rapidly quantify reads from either single-cell or bulk transcriptomes.
We downloaded the GenBank AF157706 reference transcriptome and created a kallisto index using the default -k 31 (kmer) parameter.
The best place for reproducing our HHV-6 pseudoalignment pipeline is here.
For single-cell libraries, raw sequencing reads were processed using the kallisto bus command with appropriate hyperparameters for each version
of the single-cell chemistry (either 14 or 16 bp sequence barcode and 10 or 12 bases of UMI sequence).
After barcode and UMI correction, a plain text sparse matrix was emitted, corresponding to unique HHV-6B reads
mapping to individual cells in the single-cell sequencing library. For bulk libraries, the same index could be utilized with the standard kallisto quant execution.